A flexible class of latent variable models for the analysis of antibody response data
University of Birmingham
Lund University
Presentation Overview
- A brief overview of the serology
- Current approaches for the analysis of antibody data
- The proposed modelling approach
- Application to malaria sero-epidemiology
- Summary
Key definitions
Antigen
A molecule (often a protein or polysaccharide) from a pathogen that is recognized by the immune system and can trigger an immune response.Antibody
A protein produced by B cells that specifically binds to an antigen, reflecting current or past exposure to the pathogen.
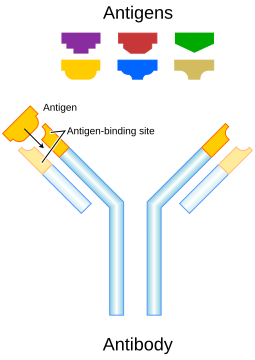
Immune response to infectious diseases
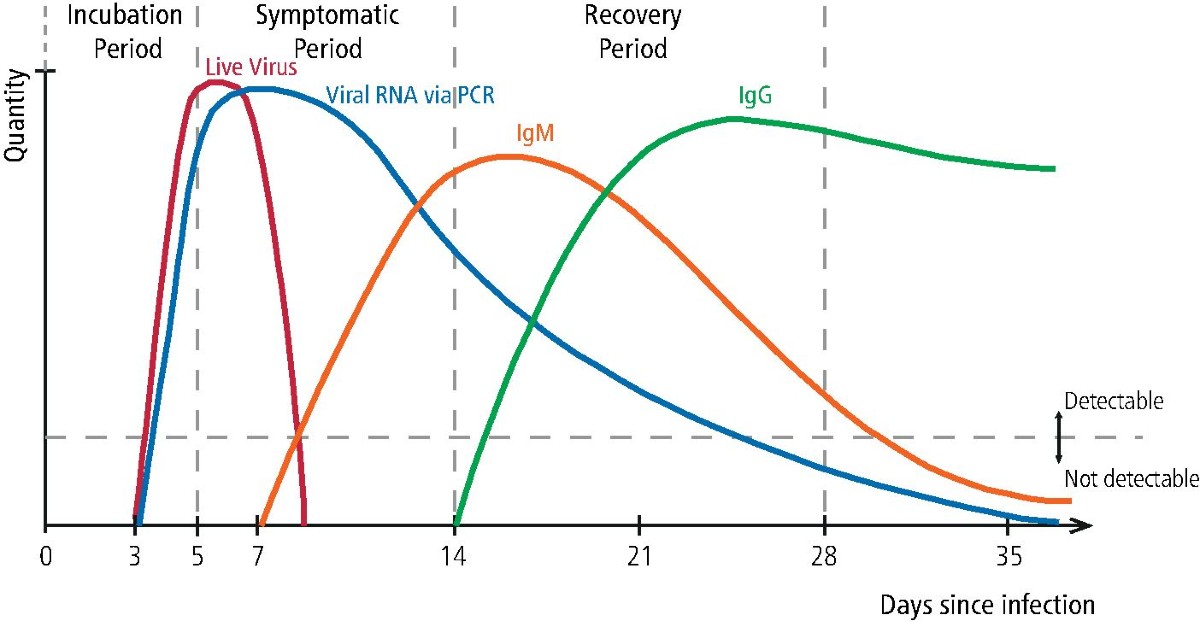
Denning DW, Kilcoyne A, Ucer C (2020). Non-infectious status indicated by detectable IgG antibody to SARS-CoV-2.
British Dental Journal, 229:521–524. Link to article
When are IgG responses informative?
Infections with many asymptomatic cases
malaria, dengue, chikungunya, ZikaAcute infections with short diagnostic windows
SARS-CoV-2, influenza, yellow feverDiseases with repeated or cumulative exposure
malaria, schistosomiasis, soil-transmitted helminths, onchocerciasisChronic infection or elimination settings
trachoma, lymphatic filariasis, onchocerciasisNot informative when cell-mediated immunity is dominant
tuberculosis, leishmaniasis
Seropositivity and seronegativity
Seronegativity
Absence of detectable antibody response to a given antigen.Seropositivity
Antibody concentration exceeding a predefined assay-specific threshold, interpreted as evidence of prior exposure or immunity.
Common approaches to serostatus classification
- Manufacturer-defined assay cut-offs
- Thresholds based on negative controls
(e.g. mean \(+\) 2 or 3 SDs)
- ROC-based cut-offs using known positive/negative samples
- Immunological correlates of protection (when available)
- Data-driven thresholds via finite mixture models
Antibody data and mixtures
- Quantitative antibody concentration \(Y_i\) (e.g. OD values)
- Classical approach: two-component Gaussian mixture
- Seronegative vs seropositive components
\[ f(y) = \pi_0 \mathcal{N}(y ; \mu_0, \sigma_0^2) + \pi_1 \mathcal{N}(y ; \mu_1, \sigma_1^2), \quad \pi_0 + \pi_1 = 1 \]
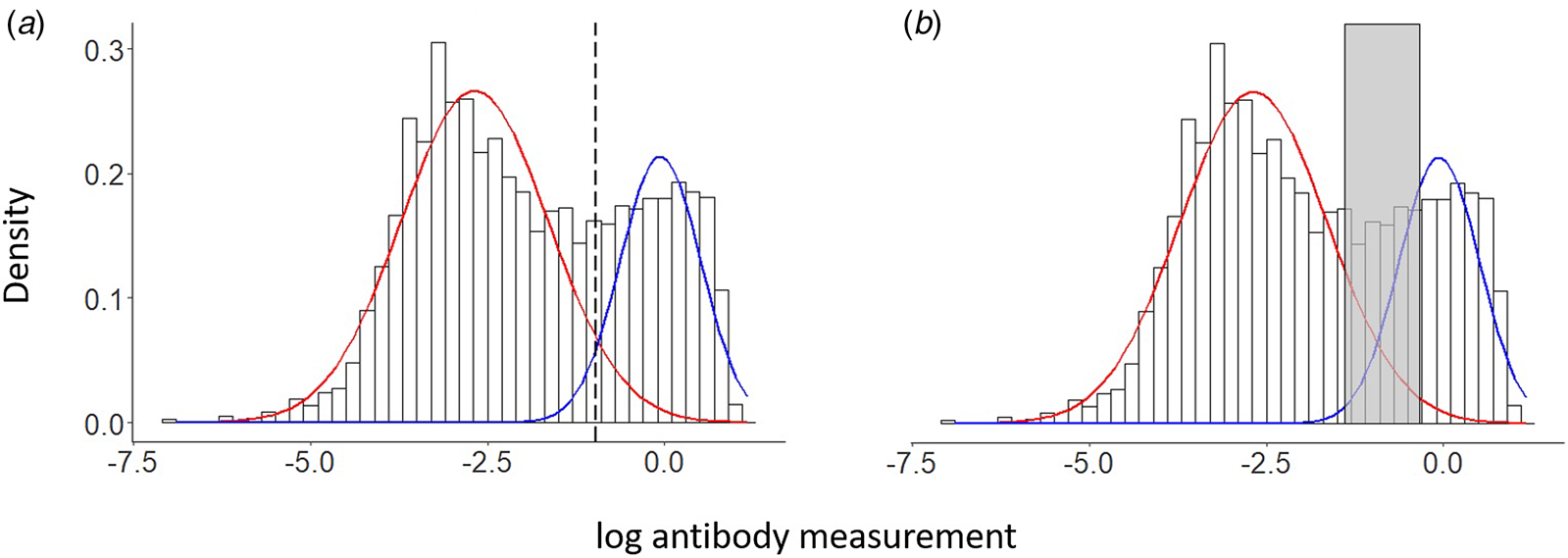
A latent variable of sero-reactivity
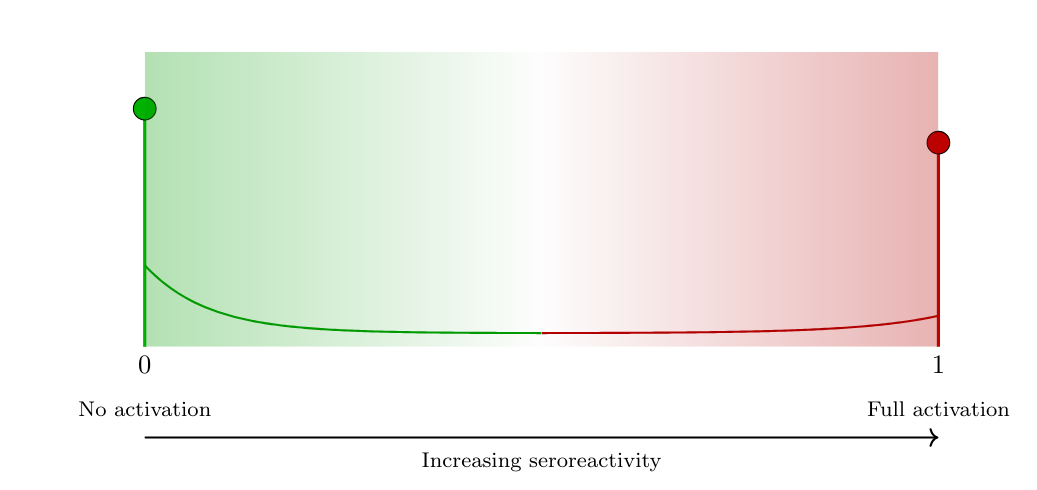
- Sero-reactivity
The presence or degree of antibody reactivity, irrespective of any diagnostic threshold.
- Latent variable \(T \in [0,1]\) represents the underlying serological activation state:
- \(T=0\): minimal or absent serological activity
- \(T=1\): strong or saturated antibody response
- \(T=0\): minimal or absent serological activity
- Antibody level given latent seroreactivity: \[
Y \mid T=t \sim
\mathcal{N}\!\big( (1-t)\mu_0 + t\mu_1,\;
(1-t)\sigma_0^2 + t\sigma_1^2 \big)
\]
- \(\mu_0\): baseline (no activation)
- \(\mu_1\): saturation level
- \(\sigma_0^2, \sigma_1^2\): heterogeneity at the two extremes
- \(\mu_0\): baseline (no activation)
Link with antibody acquisition models
Classical antibody acquisition model
(Yman et al., 2016) \[ \mathbb{E}[Y;a] = f(a) = \mu_0 + (\mu_1 - \mu_0)\{1 - \exp(-r a)\}, \] with age \(a\) and acquisition rate \(r > 0\).Interpretation in the latent sero-reactivity framework \[ \mathbb{E}[T;a] = \frac{f(a)-\mu_0}{\mu_1-\mu_0} = 1 - \exp(-r a), \]
Resulting expectation of the antibody level \[ \mathbb{E}[Y;a] = \mu_0 + (\mu_1-\mu_0)\,\mathbb{E}[T;a]. \]
Alternative latent model formulations (1)
- Model: \(Y \mid T=t = (1-t)Y_0 + tY_1\),
\(Y_0 \sim \mathcal{N}(\mu_0,\sigma_0^2),\; Y_1 \sim \mathcal{N}(\mu_1,\sigma_1^2)\) - Mean: \(\mathbb{E}(Y\mid T=t)=(1-t)\mu_0+t\mu_1\)
- Variance: \({\rm Var}(Y\mid T=t)=(1-t)^2\sigma_0^2+t^2\sigma_1^2\)
- Quadratic; minimum at \(t^*=\sigma_0^2/(\sigma_0^2+\sigma_1^2)\) → U‑shape
- Implications:
- Lowest variability at intermediate \(t\) (implausible for serology)
- Retains two latent outcome mechanisms → binary flavor
- With \(\sigma_1^2<\sigma_0^2\), our linear interpolation gives monotone decreasing variance
Alternative latent model formulations (2)
- Model: \(Y \mid T=t=(1-Z_t)Y_0+Z_tY_1,\; Z_t\sim{\rm Bernoulli}(t)\)
- Variance: \[
{\rm Var}(Y\mid T=t)=(1-t)\sigma_0^2+t\sigma_1^2+t(1-t)(\mu_1-\mu_0)^2
\]
- Extra between‑component term, maximized at \(t=0.5\)
- Implications:
- Equivalent to a GMM conditional on \(t\)
- Probabilistic binary assignment (low/high) persists
- Does not capture a continuous immune activation spectrum
How do we model the latent variable \(T\) ?
- Single-density approach
The latent sero-reactivity \(T \in [0,1]\) is modelled using a single parametric distribution: \[ T \sim \text{Beta}(\alpha,\beta), \] where \((\alpha,\beta)\) control the mean level of sero-reactivity and its heterogeneity. - Mixture-distribution approach
The latent variable \(T\) is modelled as a three-component mixture capturing graded serological states: \[ T \sim \pi_0\,\text{Beta}(\alpha_0,\beta_0) + \pi_1\,\text{Beta}(\alpha_1,\beta_1) + \pi_2\,\text{Beta}(\alpha_2,\beta_2), \] with \(\pi_k \ge 0\), \(\sum_{k=0}^2 \pi_k = 1\), representing low, intermediate, and high sero-reactivity.
Single Beta model for \(T\)
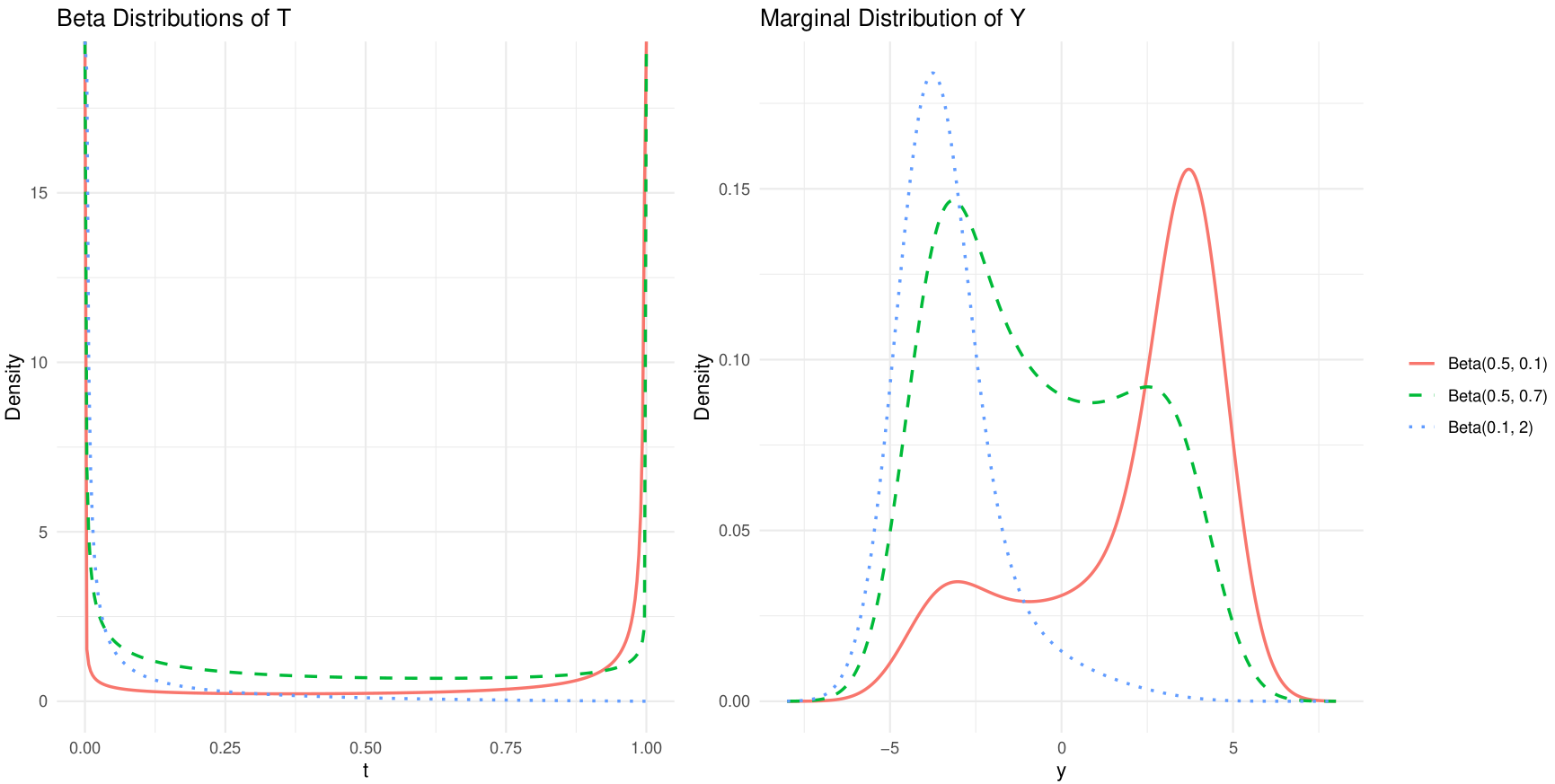
Inference
Two complementary inference approaches are used:
- full maximum likelihood, and
- a fast histogram-based approximation for exploratory fitting and initialisation.
- full maximum likelihood, and
Conditional on the latent immune state \(T\), antibody concentrations are Gaussian with mean and variance interpolating between low and high sero-reactivity extremes.
The marginal density of \(Y\) is obtained by integrating out \(T\): \[ f(y;\boldsymbol\theta,\boldsymbol\psi)=\int_0^1 \phi\!\left(y;(1-t)\mu_0+t\mu_1,(1-t)\sigma_0^2+t\sigma_1^2\right) \,g_T(t;\boldsymbol\psi)\,dt. \]Exact maximum likelihood is based on \[ \ell(\boldsymbol\theta,\boldsymbol\psi)=\sum_{i=1}^n\log f(y_i;\boldsymbol\theta,\boldsymbol\psi), \] but direct maximisation is computationally intensive due to repeated numerical integration.
A computationally efficient \(L_2\)-based estimator
To reduce computation, data are summarised into a histogram.
Let \(\widehat f_j = n_j/(n\Delta_j)\) be the empirical density in bin \(j\), and approximate model probabilities by evaluation at bin midpoints: \[ p_j(\boldsymbol\theta,\boldsymbol\psi)\approx f(m_j;\boldsymbol\theta,\boldsymbol\psi)\Delta_j. \]Parameters are estimated by minimising an \(L_2\) distance between empirical and model densities: \[ Q(\boldsymbol\theta,\boldsymbol\psi)=\sum_{j=1}^J\{\widehat f_j-f(m_j;\boldsymbol\theta,\boldsymbol\psi)\}^2. \] This yields a robust minimum-distance estimator and is computationally efficient when \(J\ll n\).
Theorem. Under regularity conditions, the \(L_2\)-based estimator converges in probability to the true \((\boldsymbol\theta, \boldsymbol\psi)\).
How do we model age dependency in \(T\)?
- How should latent sero-reactivity \(T\) change with age, reflecting cumulative exposure and immune maturation?
- Age-dependent structure
Let the distribution of \(T\) depend on age \(a\) through parameters or mixing proportions, allowing gradual acquisition. - Application to malaria
Different antigens exhibit distinct age profiles of acquisition and boosting.- AMA1
Rapid acquisition in early childhood, with sero-reactivity increasing quickly at young ages. - MSP1
Slower and more gradual age-related increases, reflecting different exposure or immune dynamics.
- AMA1
AMA1: Model formulation (1)
Latent variable for age \(<\tau\):
Mixed discrete–continuous
\[ f_T(t;a)= \begin{cases} 1-\pi(a), & t=0,\\[4pt] \pi(a)\,\alpha_2\,t^{\alpha_2-1}, & 0<t<1, \end{cases} \]Probability of high sero-reactivity
\[ \pi(a)=1-\exp(-\lambda a) \]
AMA1: Model formulation (2)
Latent variable for age \(\ge\tau\):
Latent variable distribution \[ T\sim \mathrm{Beta}(\mu(a)\phi,\,[1-\mu(a)]\phi) \]
Logit–linear regression
\[ \mathrm{logit}\{\mu(a)\}=\eta_0+\eta_1\log(a) \]Continuity constraint at the change-point \(\tau\)
\[ \eta_0=\mathrm{logit}(\mu_{\tau^-})-\eta_1\log(\tau) \]Mean of \(T\) for \(a < \tau\):
\[ \mu_{\tau^-} =p_0 e^{-\tau\lambda}\frac{\alpha_1}{\alpha_1+\beta_1} +\bigl(1-p_0 e^{-\tau\lambda}\bigr) \frac{\alpha_2}{\alpha_2+\beta_2} \]
AMA1: Parameter estimates
| Parameter | Estimate | SD | 2.5% | 50% | 97.5% |
|---|---|---|---|---|---|
| \(\mu_0\) | -3.194 | 0.021 | -3.237 | -3.194 | -3.151 |
| \(\mu_1\) | 0.747 | 0.010 | 0.727 | 0.747 | 0.768 |
| \(\sigma_0\) | 0.745 | 0.013 | 0.719 | 0.745 | 0.772 |
| \(\sigma_1\) | 0.091 | 0.013 | 0.062 | 0.091 | 0.117 |
| \(\tau\) | 20.842 | 0.420 | 20.003 | 20.876 | 20.998 |
| \(\alpha_2\) | 1.498 | 0.033 | 1.436 | 1.499 | 1.577 |
| \(\lambda\) | 0.148 | 0.005 | 0.140 | 0.148 | 0.158 |
| \(\phi\) | 4.544 | 0.131 | 4.298 | 4.551 | 4.828 |
| \(\eta_1\) | -0.138 | 0.027 | -0.191 | -0.135 | -0.080 |
AMA1 analysis: model-based histograms
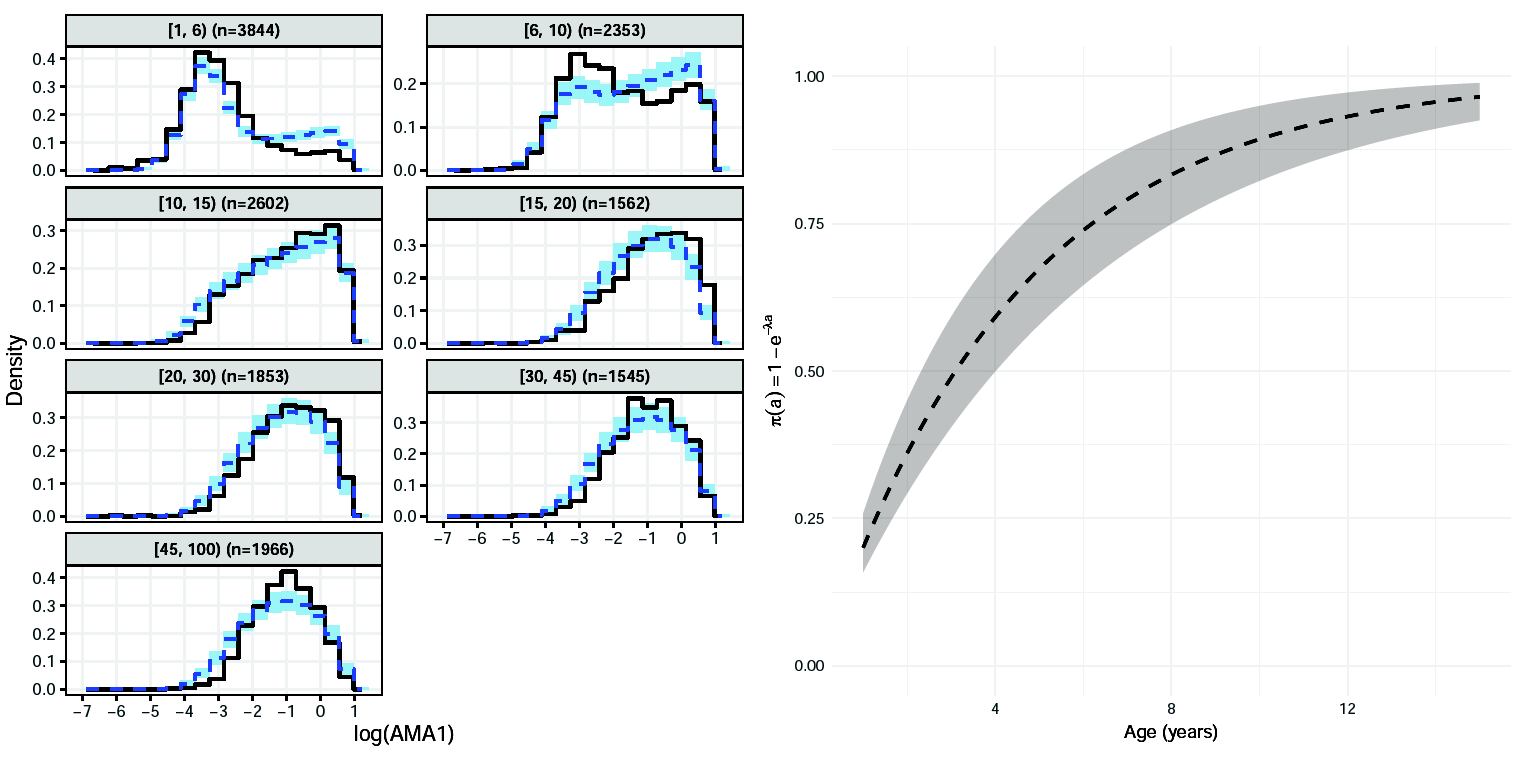
MSP1: Model formulation
Single‑component age‑dependent Beta distribution
Distribution of the latent variable \[ T \sim \mathrm{Beta}(\alpha(a),\,\beta(a)) \] where \(\alpha(a)=\alpha_0\,a^{\gamma}\) and \(\beta(a)=\beta_0\,a^{\delta(a)}\).
Change point parameter \[ \delta(a)= \begin{cases} \delta_1, & a\le \tau_{cp},\\[4pt] \delta_1+\delta_2, & a>\tau_{cp}. \end{cases} \]
MSP1: Parameter estimates
| Parameter | Mean | SD | 2.5% | 50% | 97.5% |
|---|---|---|---|---|---|
| \(\mu_0\) | -4.481 | 0.042 | -4.567 | -4.479 | -4.404 |
| \(\mu_1\) | 1.255 | 0.026 | 1.205 | 1.255 | 1.307 |
| \(\log\sigma_0\) | -0.677 | 0.043 | -0.764 | -0.676 | -0.599 |
| \(\log\sigma_1\) | -5.716 | 0.536 | -6.892 | -5.666 | -4.874 |
| \(\alpha_0\) | 0.093 | 0.036 | 0.025 | 0.093 | 0.166 |
| \(\gamma\) | 0.277 | 0.011 | 0.256 | 0.277 | 0.297 |
| \(\beta_0\) | 0.755 | 0.037 | 0.684 | 0.754 | 0.827 |
| \(\delta_1\) | 0.110 | 0.018 | 0.075 | 0.110 | 0.145 |
| \(\tau_{cp}\) | 11.623 | 0.406 | 11.004 | 11.667 | 12.197 |
| \(\delta_2\) | -0.061 | 0.010 | -0.080 | -0.061 | -0.042 |
MSP1 analysis: results
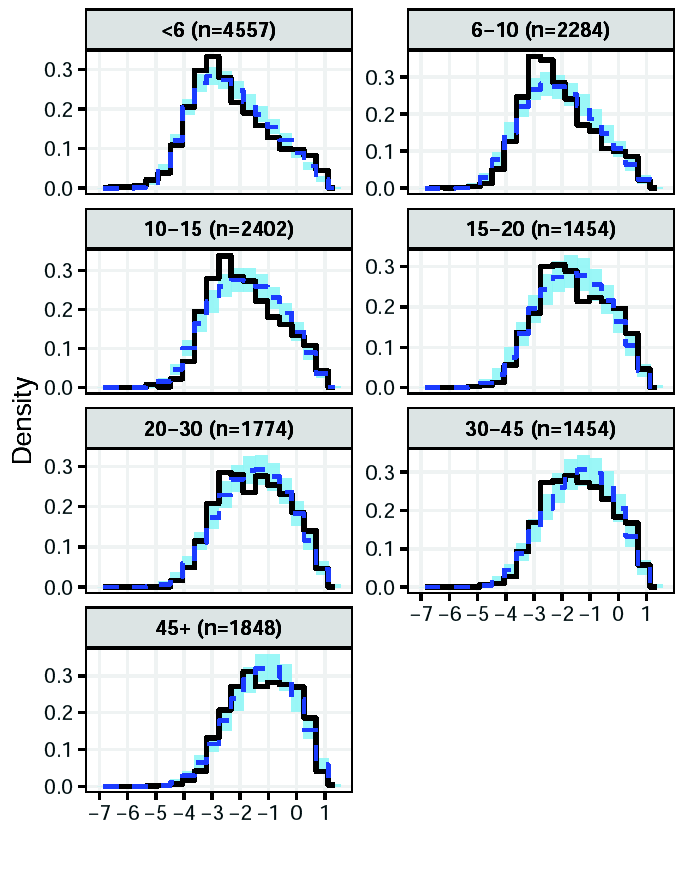
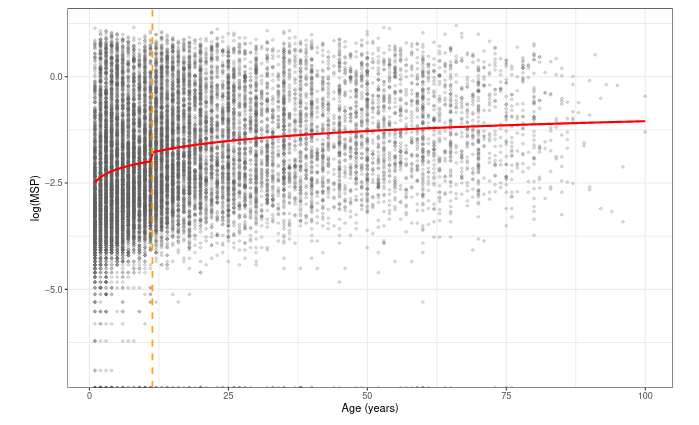
Predicted antibody levels in AMA1
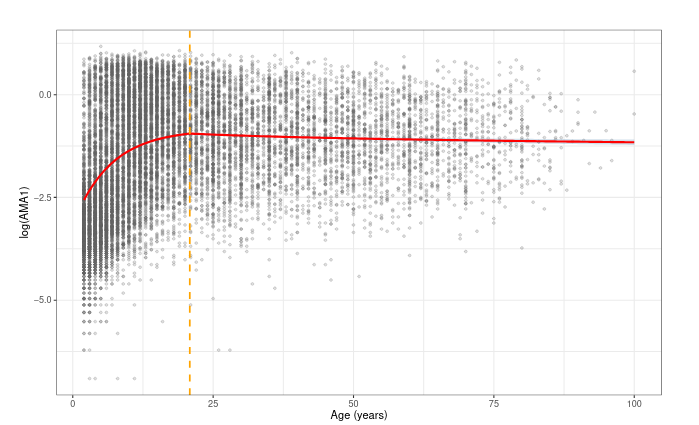
Predicted antibody levels in MSP1
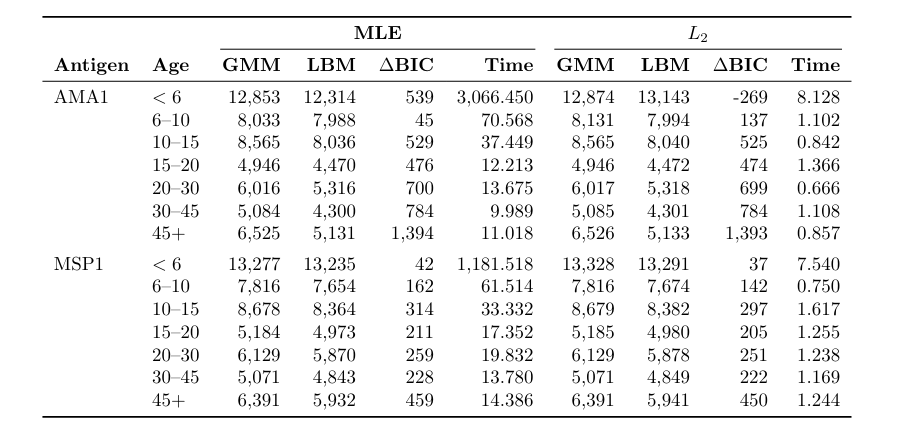
Model comparison and computational cost
Summary and conclusions
- We proposed a latent-variable framework that generalizes finite mixture models.
- It allows flexible age-dependent modelling.
- Modelling of the latent variable (seroreactivity) can be done using data-driven or mechanistic approaches.
- We are working to extend the proposed model to
- analyse geostatistical and longitudinal data-sets;
- jointly model multiple antibodies.
THANK YOU!

🔗 giorgistat.github.io
📧 e.giorgi@bham.ac.uk
📍 BESTEAM, Department of Applied Health Sciences, University of Birmingham
Exploratory analysis: AMA1
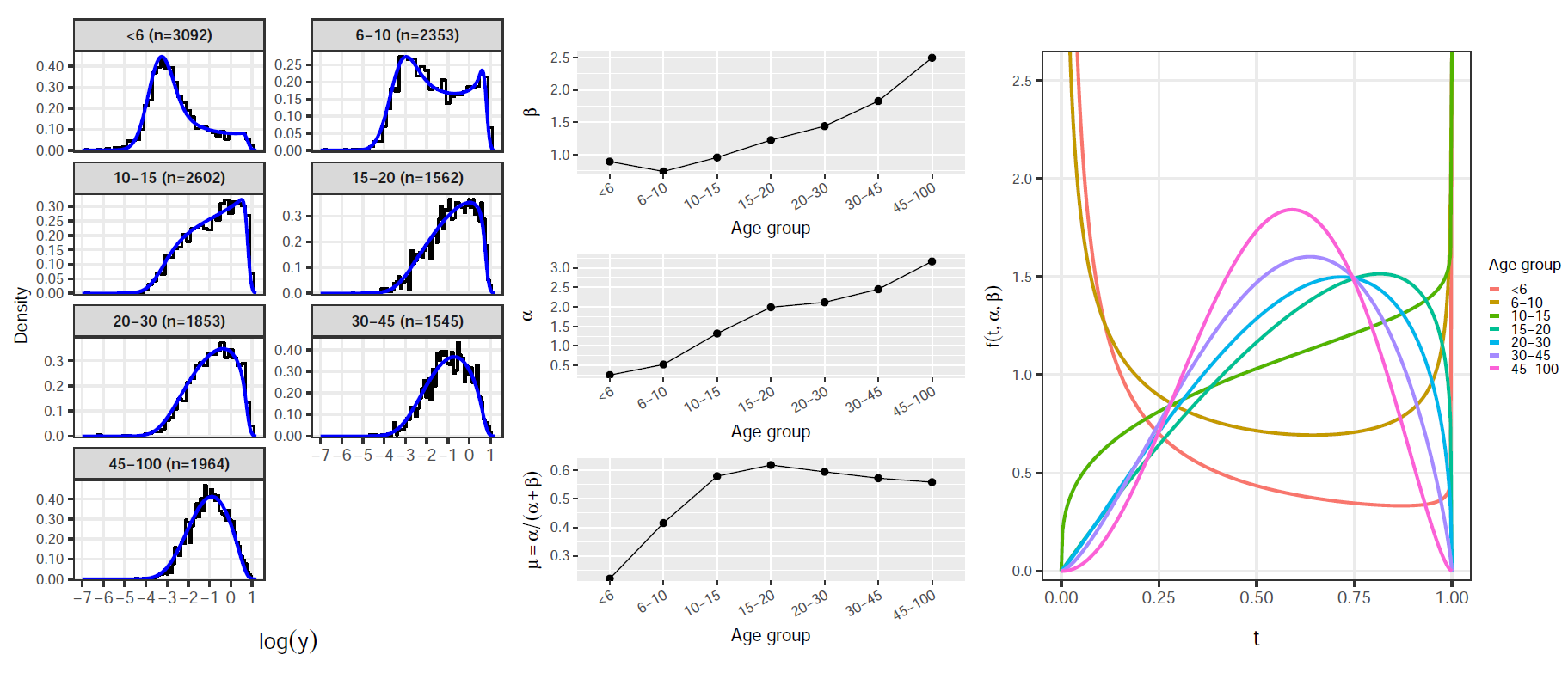
Exploratory analysis: MSP1
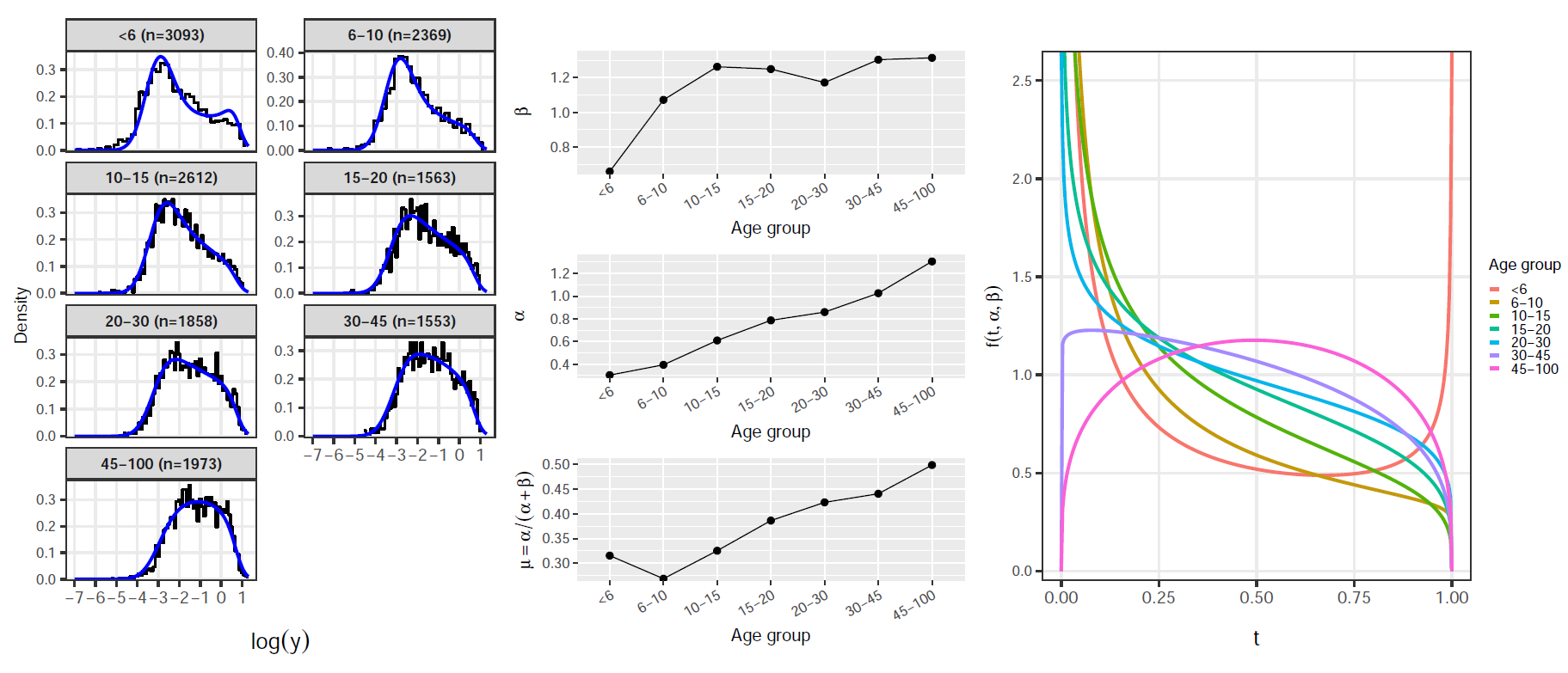
Antibody measurement: How ELISA Works
